SOM cluster: 1104
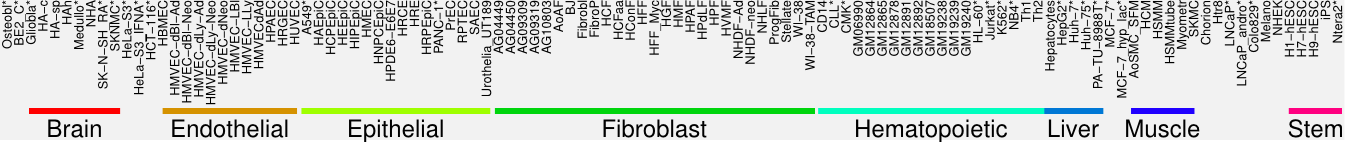
Cluster Hypersensitivity Profile
Genomic Location Trend
These plots show the distribution of the DHS sites surrounding the Transcript Start Site of the nearest gene.
Stats
Number of sites: 439Promoter: 7%
CpG-Island: 6%
Conserved: 28%
Enriched Motifs & Matches
Match Detail: [Jaspar]

| ||
|---|---|---|
| Sites: 66/100 | e-val: 2.2e-30 | ||
| Factor | e-val(match) | DB |
| TFAP2A | 0.0000014169 | JASPAR |
| SP1 | 0.000026923 | JASPAR |
| INSM1 | 0.000051268 | JASPAR |
| EBF1 | 0.00038502 | JASPAR |
| Klf4 | 0.00040063 | JASPAR |

| ||
| Sites: 68/100 | e-val: 3.7e-16 | ||
| Factor | e-val(match) | DB |
| TFAP2A | 0.000020126 | JASPAR |
| SP1 | 0.0023659 | JASPAR |
| ESR1 | 0.0039106 | JASPAR |
| Pax4 | 0.0045581 | JASPAR |
| RREB1 | 0.0049132 | JASPAR |

| ||
| Sites: 29/100 | e-val: 0.0033 | ||
| Factor | e-val(match) | DB |
| Stat3 | 0.00011191 | JASPAR |
| FEV | 0.0010937 | JASPAR |
| SPI1 | 0.0025825 | JASPAR |
| Myf | 0.0029448 | JASPAR |
| TLX1::NFIC | 0.011561 | JASPAR |

| ||
| Sites: 32/100 | e-val: 0.29 | ||
| Factor | e-val(match) | DB |
| Tal1::Gata1 | 0.0000034898 | JASPAR |
| MZF1_1-4 | 0.0077529 | JASPAR |
| PPARG::RXRA | 0.0097865 | JASPAR |
| CTCF | 0.014474 | JASPAR |
| SPIB | 0.015474 | JASPAR |
BED file downloads
Top 10 Example Regions
| Location | Gene Link | Dist. |
|---|---|---|
| chr11: 71952565-71952715 | INPPL1 | 37.82 |
| chr13: 103052240-103052390 | FGF14-IT1 | 49.96 |
| chr1: 161088980-161089130 | DEDD | 51.47 |
| chr1: 161088980-161089130 | PFDN2 | 51.47 |
| chr1: 161088980-161089130 | USP21 | 51.47 |
| chr11: 68531820-68531970 | GAL | 54.15 |
| chr5: 179599520-179599670 | RASGEF1C | 55.2 |
| chr4: 134072660-134072810 | PCDH10 | 55.34 |
| chr4: 134072660-134072810 | RP11-9G1.3 | 55.34 |
| chr22: 50962745-50962895 | NCAPH2 | 56 |