SOM cluster: 272
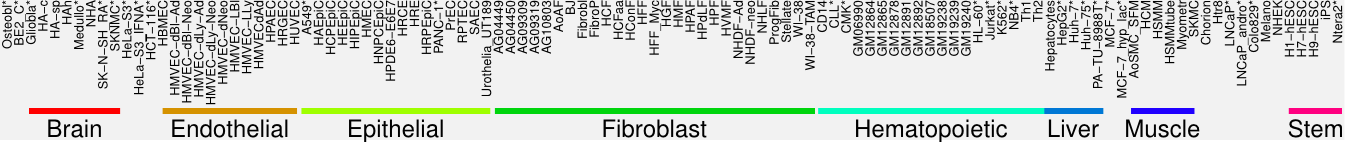
Cluster Hypersensitivity Profile
Genomic Location Trend
These plots show the distribution of the DHS sites surrounding the Transcript Start Site of the nearest gene.
Stats
Number of sites: 153Promoter: 4%
CpG-Island: 0%
Conserved: 48%
Enriched Motifs & Matches
Match Detail: [Jaspar]

| ||
|---|---|---|
| Sites: 37/100 | e-val: 2.8e-20 | ||
| Factor | e-val(match) | DB |
| AP1 | 0.00000000054528 | JASPAR |
| NFE2L2 | 0.000000017774 | JASPAR |
| NFE2L1::MafG | 0.000090024 | JASPAR |
| PBX1 | 0.0047479 | JASPAR |
| RXRA::VDR | 0.020629 | JASPAR |

| ||
| Sites: 46/100 | e-val: 0.0000000047 | ||
| Factor | e-val(match) | DB |
| RUNX1 | 0.000000021662 | JASPAR |
| ZNF354C | 0.0027538 | JASPAR |
| RREB1 | 0.0082355 | JASPAR |
| MYC::MAX | 0.043138 | JASPAR |
| Tcfcp2l1 | 0.050157 | JASPAR |

| ||
| Sites: 25/100 | e-val: 0.000072 | ||
| Factor | e-val(match) | DB |
| IRF1 | 0.00064087 | JASPAR |
| Foxd3 | 0.0018587 | JASPAR |
| FOXI1 | 0.012289 | JASPAR |
| FOXO3 | 0.014143 | JASPAR |
| Foxq1 | 0.022122 | JASPAR |
BED file downloads
Top 10 Example Regions
| Location | Gene Link | Dist. |
|---|---|---|
| chr1: 75118640-75118790 | CRYZ | 48.62 |
| chr10: 98433700-98433850 | PIK3AP1 | 51.96 |
| chr14: 54361360-54361510 | ATP5C1P1 | 56.25 |
| chr2: 46720765-46720915 | TMEM247 | 64.2 |
| chr6: 16636540-16636690 | ATXN1 | 70.18 |
| chr5: 33850300-33850450 | ADAMTS12 | 71.28 |
| chr5: 83140440-83140590 | EDIL3 | 74.9 |
| chr9: 35848205-35848355 | LINC00961 | 80.54 |
| chr9: 35848205-35848355 | FAM221B | 80.54 |
| chr7: 139424940-139425090 | HIPK2 | 81.73 |