SOM cluster: 632
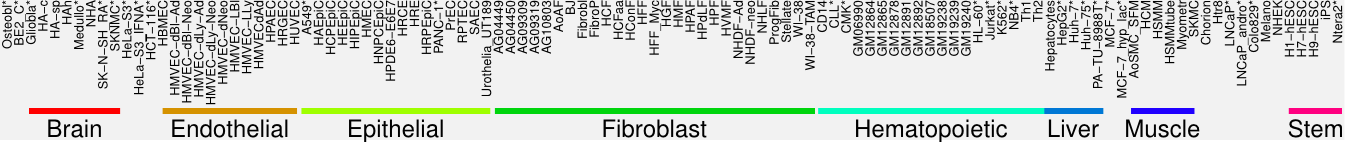
Cluster Hypersensitivity Profile
Genomic Location Trend
These plots show the distribution of the DHS sites surrounding the Transcript Start Site of the nearest gene.
Stats
Number of sites: 135Promoter: 41%
CpG-Island: 25%
Conserved: 44%
Enriched Motifs & Matches
Match Detail: [Jaspar]

| ||
|---|---|---|
| Sites: 14/100 | e-val: 0 | ||
| Factor | e-val(match) | DB |
| MEF2A | 0.00091307 | JASPAR |
| NFATC2 | 0.0081716 | JASPAR |
| FOXO3 | 0.0091713 | JASPAR |
| T | 0.02534 | JASPAR |
| NFE2L2 | 0.038313 | JASPAR |

| ||
| Sites: 14/100 | e-val: 0.000000028 | ||
| Factor | e-val(match) | DB |
| RXRA::VDR | 0.0012415 | JASPAR |
| REST | 0.0032475 | JASPAR |
| EWSR1-FLI1 | 0.0093744 | JASPAR |
| RXR::RAR_DR5 | 0.0094143 | JASPAR |
| PPARG | 0.0097702 | JASPAR |

| ||
| Sites: 11/100 | e-val: 0.28 | ||
| Factor | e-val(match) | DB |
| Egr1 | 0.031239 | JASPAR |
| MYC::MAX | 0.037956 | JASPAR |
| SPIB | 0.039031 | JASPAR |
| Myb | 0.047296 | JASPAR |
| NHLH1 | 0.075108 | JASPAR |

| ||
| Sites: 9/100 | e-val: 0.0037 | ||
| Factor | e-val(match) | DB |
| TFAP2A | 0.010754 | JASPAR |
| SP1 | 0.021194 | JASPAR |
| ESR1 | 0.023621 | JASPAR |
| TLX1::NFIC | 0.026233 | JASPAR |
| Myb | 0.033834 | JASPAR |

| ||
| Sites: 7/100 | e-val: 0.079 | ||
| Factor | e-val(match) | DB |
| Stat3 | 0.00049599 | JASPAR |
| FEV | 0.010432 | JASPAR |
| SPI1 | 0.011185 | JASPAR |
| PLAG1 | 0.032923 | JASPAR |
| RXR::RAR_DR5 | 0.070611 | JASPAR |