SOM cluster: 97
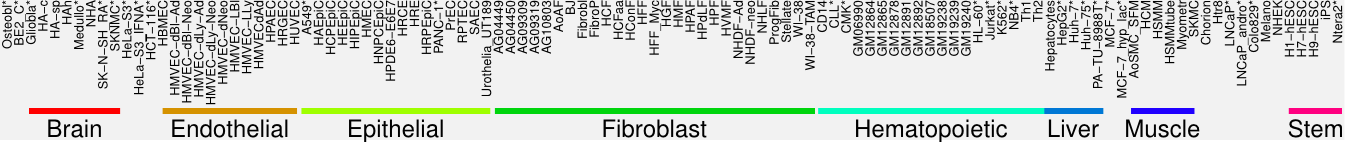
Cluster Hypersensitivity Profile
Genomic Location Trend
These plots show the distribution of the DHS sites surrounding the Transcript Start Site of the nearest gene.
Stats
Number of sites: 569Promoter: 88%
CpG-Island: 95%
Conserved: 87%
Enriched Motifs & Matches
Match Detail: [Jaspar]

| ||
|---|---|---|
| Sites: 97/100 | e-val: 0 | ||
| Factor | e-val(match) | DB |
| TFAP2A | 0.00089499 | JASPAR |
| Egr1 | 0.002197 | JASPAR |
| PLAG1 | 0.021175 | JASPAR |
| E2F1 | 0.044647 | JASPAR |
| SP1 | 0.050248 | JASPAR |

| ||
| Sites: 98/100 | e-val: 0 | ||
| Factor | e-val(match) | DB |
| SP1 | 0.000000044288 | JASPAR |
| Klf4 | 0.00010108 | JASPAR |
| TFAP2A | 0.00012137 | JASPAR |
| Egr1 | 0.0081699 | JASPAR |
| PLAG1 | 0.041818 | JASPAR |

| ||
| Sites: 69/100 | e-val: 9.5e-29 | ||
| Factor | e-val(match) | DB |
| SP1 | 0.0000000053617 | JASPAR |
| Klf4 | 0.00028791 | JASPAR |
| RREB1 | 0.00061061 | JASPAR |
| Egr1 | 0.0030787 | JASPAR |
| TFAP2A | 0.016396 | JASPAR |

| ||
| Sites: 18/100 | e-val: 0.039 | ||
| Factor | e-val(match) | DB |
| Egr1 | 0.00051096 | JASPAR |
| NHLH1 | 0.00087548 | JASPAR |
| CEBPA | 0.018824 | JASPAR |
| E2F1 | 0.028653 | JASPAR |
| TP53 | 0.046192 | JASPAR |

| ||
| Sites: 32/100 | e-val: 0.026 | ||
| Factor | e-val(match) | DB |
| SP1 | 0.000044939 | JASPAR |
| Klf4 | 0.0068424 | JASPAR |
| Egr1 | 0.025102 | JASPAR |
| PLAG1 | 0.051743 | JASPAR |
| Tal1::Gata1 | 0.052214 | JASPAR |
BED file downloads
Top 10 Example Regions
| Location | Gene Link | Dist. |
|---|---|---|
| chr12: 77459360-77459510 | E2F7 | 39.03 |
| chr9: 35829140-35829290 | FAM221B | 44.83 |
| chr17: 77070860-77071010 | ENGASE | 45.78 |
| chr1: 161129200-161129350 | PVRL4 | 51.19 |
| chr1: 161129200-161129350 | KLHDC9 | 51.19 |
| chr17: 7465140-7465290 | AC113189.5 | 52.58 |
| chr17: 7465140-7465290 | SLC35G6 | 52.58 |
| chr7: 65540640-65540790 | RP5-1132H15.2 | 53.99 |
| chr6: 31164860-31165010 | PSORS1C3 | 54.57 |
| chr6: 31164860-31165010 | WASF5P | 54.57 |